This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Delayed Sleep Phase Syndrome?
|
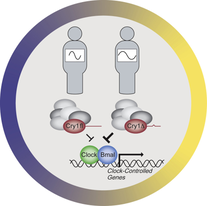
Delayed sleep phase syndrome (DSPS) is a circadian rhythm sleep disorder, in which a person has the propensity of sleeping and waking at later hours than conventional times of the day. DSPS can inhibit the ability to be well-rested and awake at desired times for work or school. Individuals with DSPS can achieve normal sleep quality and quantity when allowed to choose their schedule. Typical times for individuals with delayed sleep phase to go to bed are between 1 and 4 am, and wake up are between 8 and 11 am.
|
What is CRY1?
The CRY1 (cryptochrome circadian regulator 1) gene is located on chromosome 12 in humans and is linked to the regulation of the circadian rhythm, encoding a binding protein for the circadian core oscillator complex. CRY1 gene codes for the CRY1 protein.
Gene Ontology Terms of CRY1
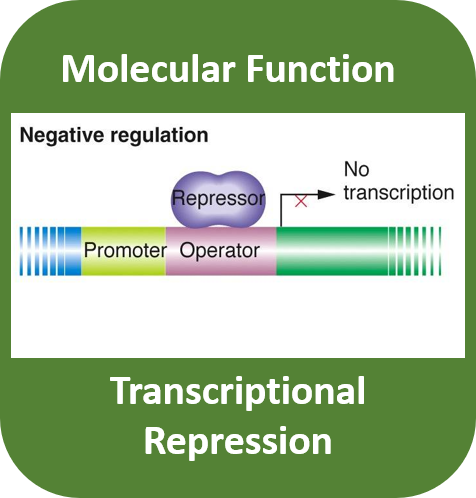
Gene ontology terms help understand the role of CRY1 in regulating the mechanisms of the circadian rhythm between and within cells. CRY1 helps coordinate the length of a circadian cycle through transcriptional repressing CLOCK and BMAL located within the nucleus of all cells.
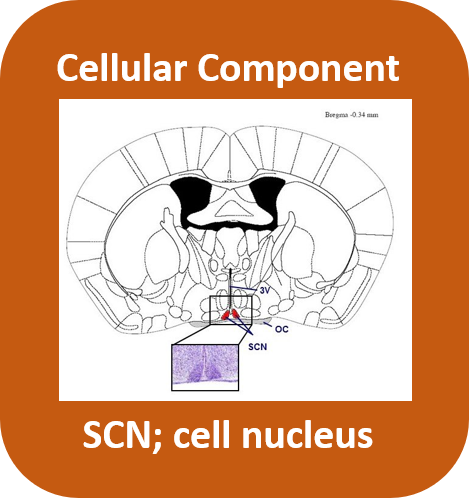
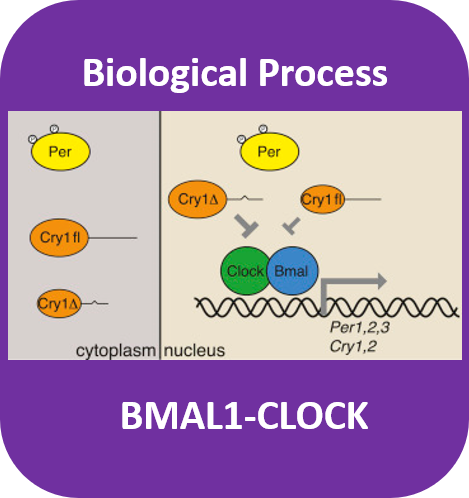
The CRY1 gene codes for the CRY1 protein containing a photolyase domain and a flavin adenin dinucleotide binding domain. This protein can be found in the nucleus of all cells in the peripheral clocks and especially at the suprachiasmatic nucleus in the hypothalamus as the central clock whose main function is to control most circadian rhythms in the body. The CRY1 protein serves as a transcriptional repressor of the feedback loop producing the transcriptional factors CLOCK and BMAL1 that allow for the circadian clocks to restart.
Gene Ontology Terms of CRY1
Gene ontology terms help understand the role of CRY1 in regulating the mechanisms of the circadian rhythm between and within cells. CRY1 helps coordinate the length of a circadian cycle through transcriptional repressing CLOCK and BMAL located within the nucleus of all cells.
The CRY1 gene codes for the CRY1 protein containing a photolyase domain and a flavin adenin dinucleotide binding domain. This protein can be found in the nucleus of all cells in the peripheral clocks and especially at the suprachiasmatic nucleus in the hypothalamus as the central clock whose main function is to control most circadian rhythms in the body. The CRY1 protein serves as a transcriptional repressor of the feedback loop producing the transcriptional factors CLOCK and BMAL1 that allow for the circadian clocks to restart.
Protein Domains of CRY1
|
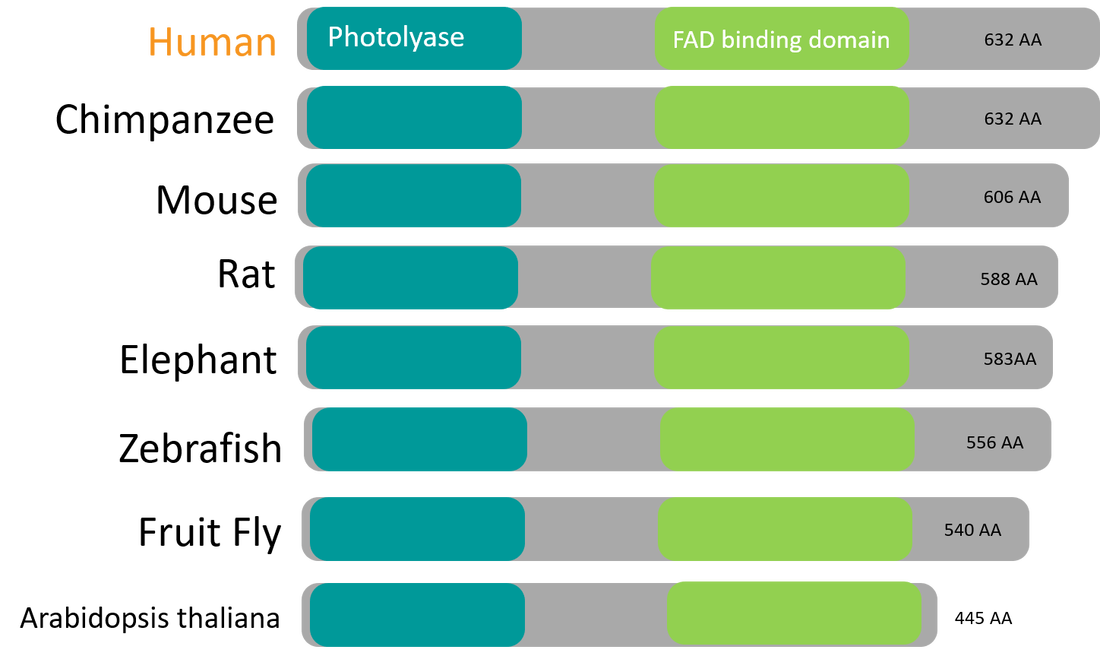
CRY1 consists of two protein domains, a photolyase domain and a flavin adenine dinucleotide domain. The characterization of CRY1 protein domains for other organisms was obtained with SMART and Pfam, demonstrating amino acid sequences that are highly conserved between mammals, invertebrates, and plants. Amino acid conservation within species provides insight with respect to the importance of the protein for proper circadian rhythm regulation.
Homology of CRY1 across species Protein homology occurs when there is shared ancestry between proteins in different species. As the percent identity between proteins increases, sequences are more similar and homology can be inferred. Protein homology of CRY1 is highly conserved across species. High percent identity allows us to choose from a multitude of organisms to study the protein and gene and correlate findings to human mechanisms and diseased states. While highest conservation occurs with mammals, other organisms like zebrafish demonstrate great similarities in function and localization of the protein allowing for efficient study of CRY1. |
Phylogeny of CRY1 protein
|
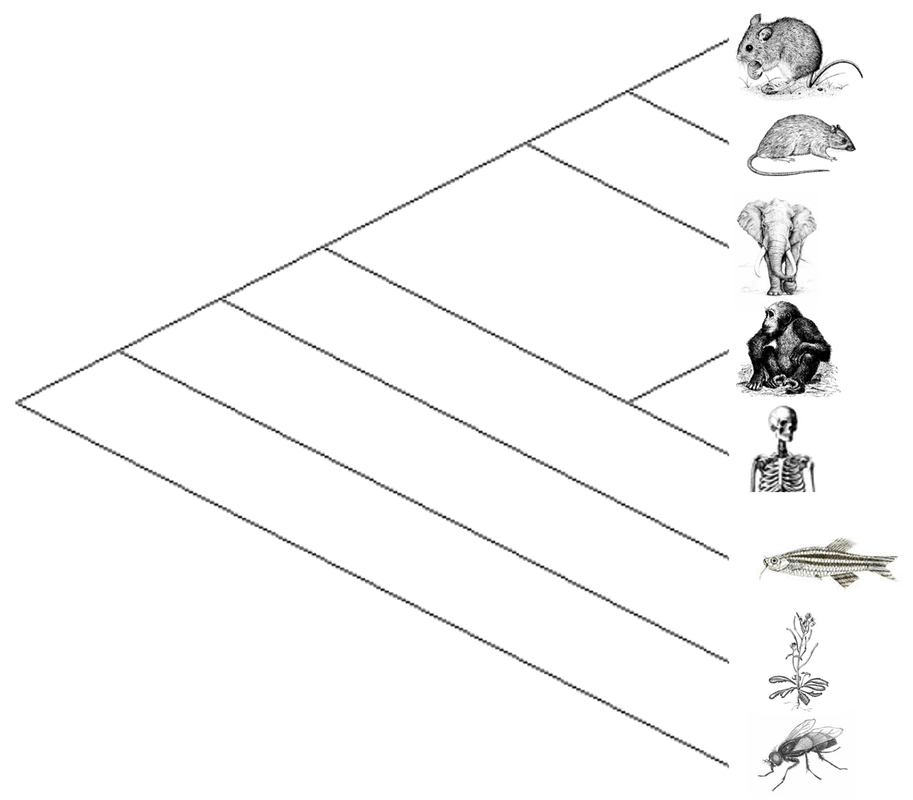
Phylogenetic trees represent the evolutionary relationships among a group of organisms based on shared derived traits coming from a most common ancestor. The comparison and analysis of a combination of external morphology, internal anatomy, behaviors, biochemical pathways, DNA and protein sequences, and fossils can be used to build phylogenetic trees.
The neighbor-joining phylogenetic tree showcases most parsimonious relationships between the organisms' protein. Proximity of zebrafish and mice to humans, make them both good candidates for studying CRY1. |
What model organism can be used to study DSPS?
|
Model organisms are useful in understanding disease and bodily mechanisms in humans, while being easy to maintain and breed or transform for specific phenotypes. Understanding the advantages of different model organisms allows for efficient studies. Due to the high conservation of Cry1 in species, different organisms can be used to study the effects of Cry1 in regulation of bodily functions in humans. Zebrafish are small, can be easily and rapidly bred or transformed for specific mutations, and their behavior can be monitored efficiently, perfect subjects for circadian rhythm and Cry1 studies.
|
What is the gap in knowledge?
|
Recent studies have encountered a mutation in the CRY1 mechanism linked to DSPS. A deletion on chromosome 12 causes a change in the transcriptional activity of CRY1 by modifying the tail region of the protein after the photolyase region. However, no studies have been performed yet to understand the mechanism influenced by the CRY1 that affects the activity of the Clock/Bmal1 transcription factors.
|
What is my primary goal?
My primary goal will be to understand the mechanisms of the CRY1 protein tail in regulating the circadian clock. First I will determine the conservation of amino acid domains between normal and mutant lines. Second I will identify chemical compounds that could offset the effects of the mutations, and finally I will determine if any changes in protein interactions occur when CRY1 is mutated.
|
Aim #1: Determine the conservation of amino acid domains of CRY1 and mutant sequences to determine effects of deleting the Cry1 protein tail.
Approach: Use Clustal Omega to align normal and wild-type sequences and demonstrate the conservation between the lines. Hypothesis: Deletions in the Cry1 gene will result in a shortening of the protein domains translated, will destabilize the protein, and will lead to the lengthening of sleep-wake cycle in the transgenic lines demonstrating the importance of this portion in the regulation of the CLOCK/BMAL1 circadian clock. Rationale: If there are changes in the protein domains in diseases states of the CRY1 protein, more focused studies will be possible to understand the function of the protein in different states. |
|
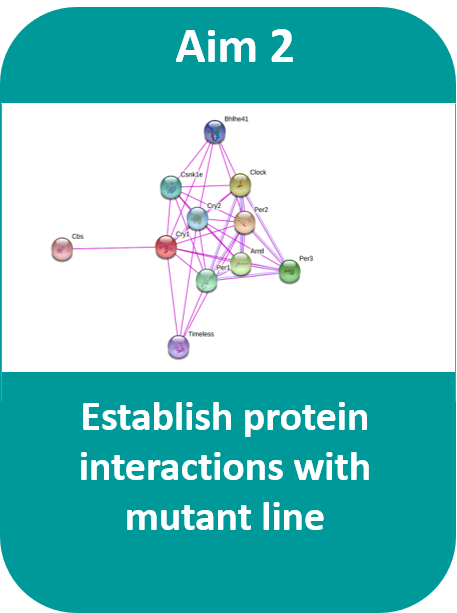
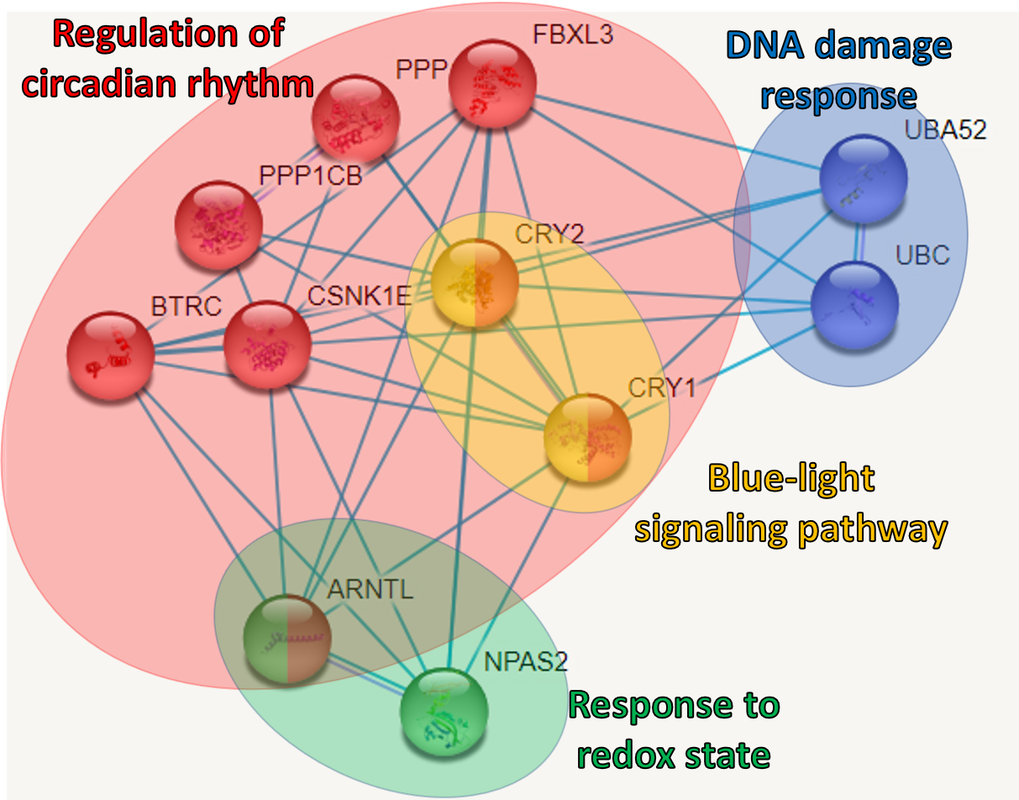
Aim #2: Establish the protein interactions of the transgenic zebrafish lines with the protein tail deletion.
Approach: Extraction of CRY1 and its interacting proteins will be characterized with mass spectrometry and bioinformatic analysis. Gene ontology terms will be used to define the changes in interactions. Hypothesis: Mutations in the Cry1 sequence will change the protein interactions required for normal modulation of circadian clock length. Rationale: Determining the changes in Cry1 protein interactions with a deletion in its tail will provide insight on the role of this segment in the circadian rhythm. By looking at the changes in protein interactions, we can learn more about the mechanisms where Cry1 takes part in. |
|
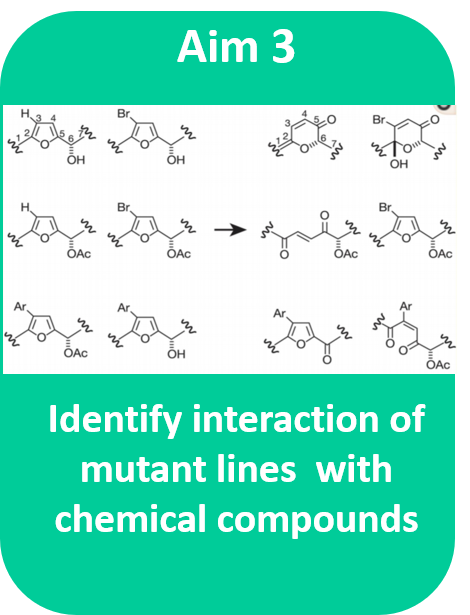
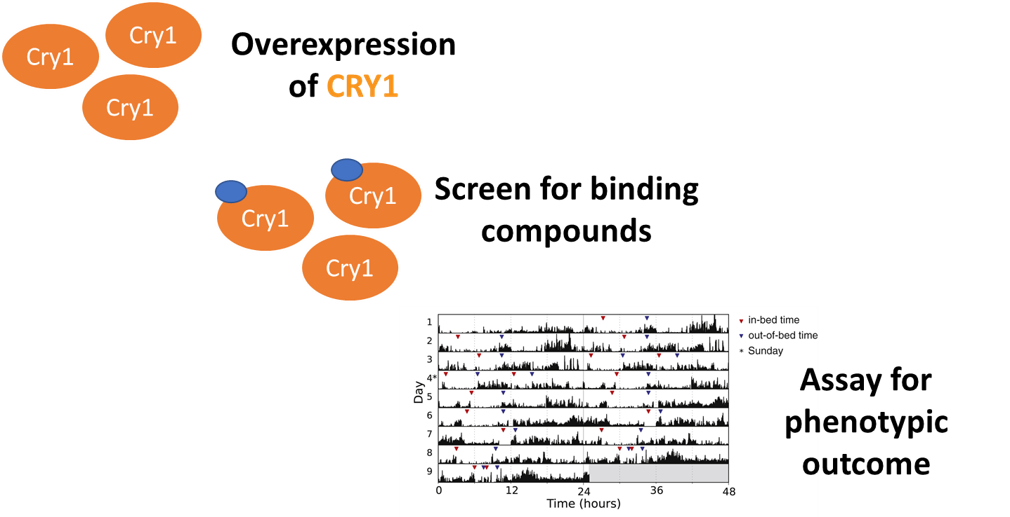
Aim #3: Identify chemical compounds that will interact with and potentially repress the effects of the Cry1 tail deletion.
Approach: Assemble a chemical library and perform chemical genomic screen on transgenic zebrafish lines to find interacting compounds changing the phenotype Hypothesis: Small molecules that can restore the phosphorylation effects of the Cry1 protein tail and will reestablish the length of the circadian rhythm of the mutagenic lines. Rationale: Compounds that restore the activity patterns of the mutagenic lines will most likely function and bind similarly to the function of the missing protein tail segment. Treatments for DSPS can be designed. |
Summary
- A deletion in CRY1 gene results in the overexpression of the protein and lengthening of the circadian cycle.
- The study of mutagenic lines allows for the study of protein interactions in diseased states.
- Understanding the mechanisms behind the CRY1 protein tail will allow for the development of therapies and treatment for circadian rhythm disorders.
- The study of mutagenic lines allows for the study of protein interactions in diseased states.
- Understanding the mechanisms behind the CRY1 protein tail will allow for the development of therapies and treatment for circadian rhythm disorders.
Future Directions
The information gathered through this website put together information about the CRY1, while opening up possibilities for further studies in the effects of this protein in circadian rhythm regulation. Understanding the mechanisms of the protein tail of CRY1 protein will help us develop treatments and strategies to help individuals with Delayed Sleep Phase Syndrome and other sleep-wake cycle disorders.
Further studies in the localization of CRY1 expression will also allow the development of focused therapies and personalization of treatment between male and females individuals.
Further studies in the localization of CRY1 expression will also allow the development of focused therapies and personalization of treatment between male and females individuals.
| acosta_finaltalk_draft1_4-5-19.pptx | |
| File Size: | 6578 kb |
| File Type: | pptx |
| acosta_finaltalk_final_4-29-19.pptx | |
| File Size: | 5870 kb |
| File Type: | pptx |
| Final Talk PDF | |
| File Size: | 741 kb |
| File Type: | |
|
Contact me
Sara Acosta Villarreal Genetics and Genomics, UW-Madison [email protected] Last updated: May 10, 2019 |
Visitors Worldwide
|