This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is transcriptomics?
Transcriptomics is the study and sequencing of the transcriptome [1]. The transcriptome is the set of all RNA molecules in one single cell of in a population of cells reflecting the expression of genes at a given time or under specific conditions.
How is the transcriptome studied?
RNA measurement and analysis can be performed with simple methods such as microarrays and reverse transcriptase-PCR, and with more technologically advanced methods like next generation sequencing systems [1]. Transcriptomics is helping researchers understand the regulation of development and of disease and other processes of interest in the human body.
Transcriptomic Analysis Methods [4]
|
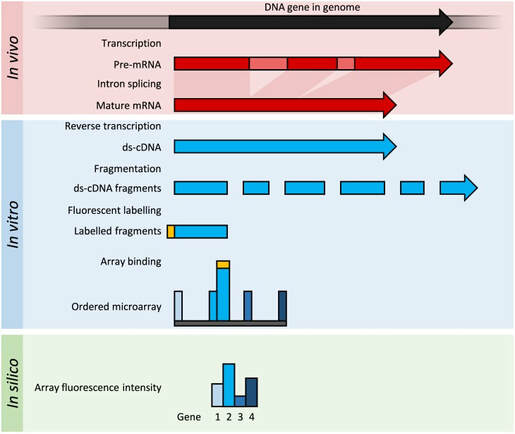
Microarrays detect nucleic acids in a sample by hybridization to probes on microchips. RNA samples of different conditions are converted into complementary DNA and fragmented. The fragments are then tagged with fluorescent tags and are hybridized into microchips with known transcript probes.
|
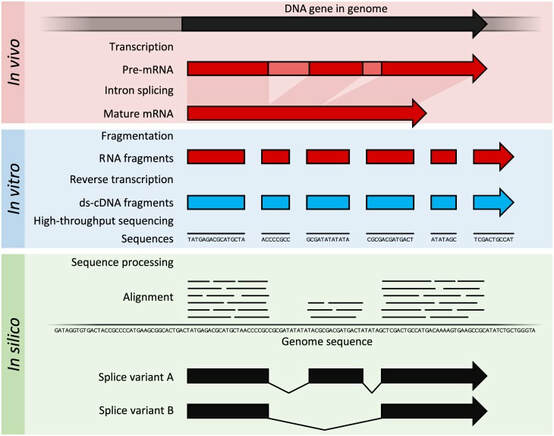
RNA-Sequencing is a next-generation sequencing method that starts with the extraction of all RNA in samples of different conditions. Not unlike microarrays, the RNA extracted is converted into cDNA and fragmented. These fragments are then sequenced and aligned to the genome of origin. The number of fragments present will conclude the level of expression of specific gene sequences.
|
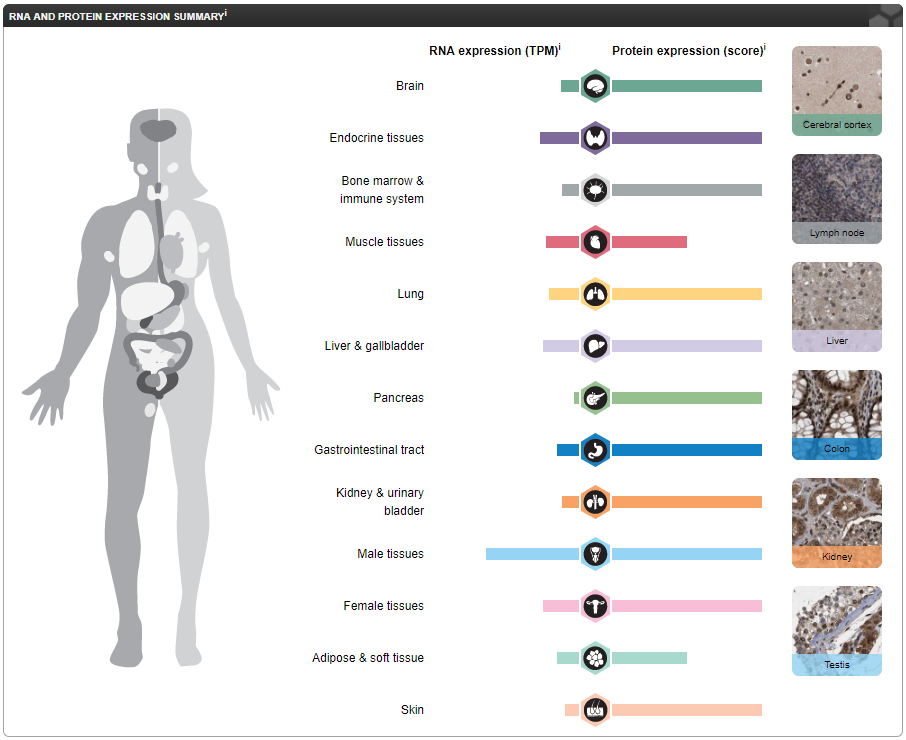
Where is CRY1 RNA expressed?
Conclusion
CRY1 is a core component of the circadian clock. Protein expression is ubiquitous within the cytoplasm and nucleus of all tissues. We see differential expression of RNA with greatest expression in male tissues (testis). While there are no reports of phenotypes for CRY1 mutations seen at different rates based on sex, this expression provides a source for further study to understand the function of CRY1 in the specific tissue type.
References.
1. What is transcriptomics? (n.d.). Retrieved from http://www.phgfoundation.org/infographic/what-is-transcriptomics
2. Introduction to Single‐Cell RNA Sequencing - Olsen - 2018 - Current Protocols in Molecular Biology - Wiley Online Library. (2019). doi:10.1002/cpmb.57
3. Papalexi, E., & Satija, R. (2017). Single-cell RNA sequencing to explore immune cell heterogeneity. Nature Reviews Immunology, 18(1), 35. doi:doi:10.1038/nri.2017.76
4. Lowe, R., Shirley, N., Bleackley, M., Dolan, S., & Shafee, T. (2017). Transcriptomics technologies. PLOS Computational Biology, 13(5). doi:10.1371/journal.pcbi.1005457
Header image.
1. What is transcriptomics? (n.d.). Retrieved from http://www.phgfoundation.org/infographic/what-is-transcriptomics
2. Introduction to Single‐Cell RNA Sequencing - Olsen - 2018 - Current Protocols in Molecular Biology - Wiley Online Library. (2019). doi:10.1002/cpmb.57
3. Papalexi, E., & Satija, R. (2017). Single-cell RNA sequencing to explore immune cell heterogeneity. Nature Reviews Immunology, 18(1), 35. doi:doi:10.1038/nri.2017.76
4. Lowe, R., Shirley, N., Bleackley, M., Dolan, S., & Shafee, T. (2017). Transcriptomics technologies. PLOS Computational Biology, 13(5). doi:10.1371/journal.pcbi.1005457
Header image.
|
Contact me
Sara Acosta Villarreal Genetics and Genomics, UW-Madison [email protected] Last updated: May 10, 2019 |
Visitors Worldwide
|